Lecture Details[]
Wayne Sturrock; Week 4 MED1022; Physiology
Lecture Content[]
Muscle fibres shorten and generate force, contraction involves actin and myosin controlled by intracellular Ca. Cardiac muscle is branched and striated with 1 central nucleus. Epimysium surrounds muscle, perimysium surrounds bundles, endomysium surrounds fibres. Whole muscle is made of muscle fibres grouped into fascicles. Single muscle cell is 10 to 100um in diameter, short (stapedius) to long (sartorius), myofibrils are cylindrical elements made up of myofilaments, myofilaments can be thick or thin and are arranged into sarcomeres.
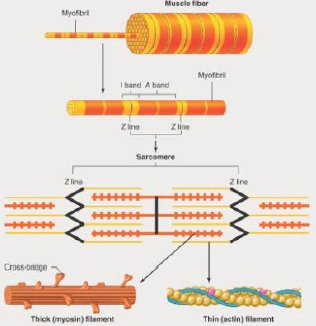
Sarcomeres are the basic contractile unit, responsible for the striated appearance. Each thick filament is surrounded by three thin filaments. A band is between two Z discs, I band is only thin filaments, A band is entire length of thick filament, H zone is central region of A band (thick filaments only) and M line is the attachment site for thick filaments, divides the A band in half.
Thin filaments: actin is a globular molecule in a 2 stranded helix. Each molecule has a binding site for myosin. Tropomyosin lies along actin filaments and covers the myosin binding sites on actin in relaxed muscle. It is part of the system for regulating the turning on and off of contraction. Troponin has TnI, T
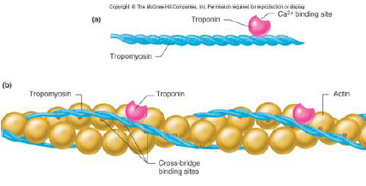
and C. It is located on regular intervals on the actin molecule and binds to both tropomyosin and actin. It binds Ca to switch contraction on. Thin filaments attach to the Z line. Myosin is a large molecule with a long, rigid tail and a globular head. They line up tail-to-tail to form thick filaments. The tails form the backbone of the thick filament and the globular heads are the crossbridge projections. Each globular head has an actin binding site and an ATPase site (to split ATP). The crossbridges rotate and pull on the thin filament, there is sliding of the filaments past each other. They pull Z lines together allowing the sarcomere to shorten- the I band and H zone shorten but the A band stays the same. The sliding requires energy, there are 4 basic steps.
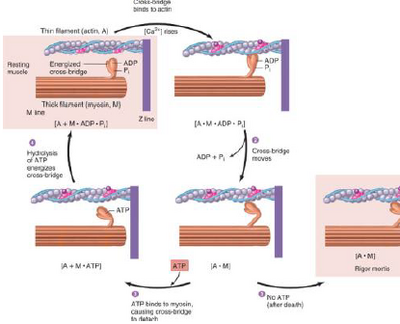
1. ATP splitting: energy is 'stored' in the crossbridge, M.ATP -> M*ADP.Pi (M* is energised myosin crossbridge)
2. Attachment (actin binding), A + M*.ADP.Pi -> A.M.ADP.Pi
3. Power stroke (product release and crossbridge rotation), A.M.ADP.Pi -> A.M + ADP + Pi
4. Detachment (fresh ATP binds to myosin) A.M + ATP -> A + M.ATP
ATP splitting primes the crossbridge to bind to actin, like setting a mouse trap. ATP is necessary to bind to myosin to let go of the actin. Each cycle moves it about 10nm. Is repeated many times during a contraction. The rate of contraction is determined by rate of splitting ATP, myosin ATPase activity is determinant of muscle speed.
In absence of Ca (below 10^7) intracellularly (in cytoplasm) tropomyosin is bound to myosin so actin can't bind. In presence (about 10^5) Ca binds to troponin which rotates pulling tropomyosin with it, exposing myosin binding sites allowing myosin to bind to actin, continues as long as Ca is around.